X-Ray Crystallography for Hit Confirmation
In the X-ray crystallography experiment, the crystalline atoms of target produce a bunch of X-rays diffract into many specific directions and a crystallographer is able to generate a three-dimensional picture of the density of electrons within the crystal. Based on this electron density, the mean positions of the atoms in the crystal can be determined and other detailed information such as their chemical bonds, disorder can also be obtained.
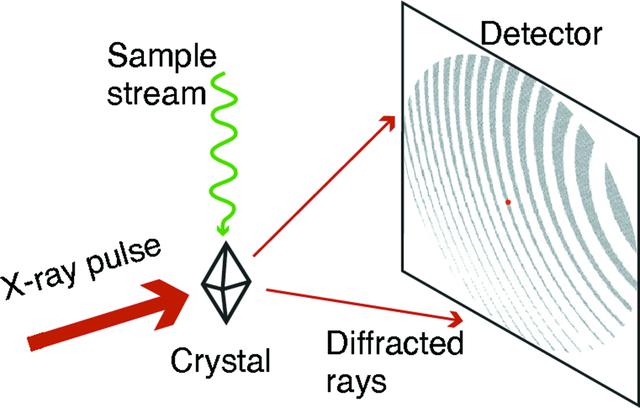 Fig.1 Generic scheme depicting the detection process in an X-ray crystallography experiment. (Ke, T. W.; et al. 2018)
Fig.1 Generic scheme depicting the detection process in an X-ray crystallography experiment. (Ke, T. W.; et al. 2018)Application of X-Ray Crystallography in Fragment-Based Screening
X-ray crystallography method can be employed in the fragment library screening and it is also applied as the final approach to confirm the predictive hit.
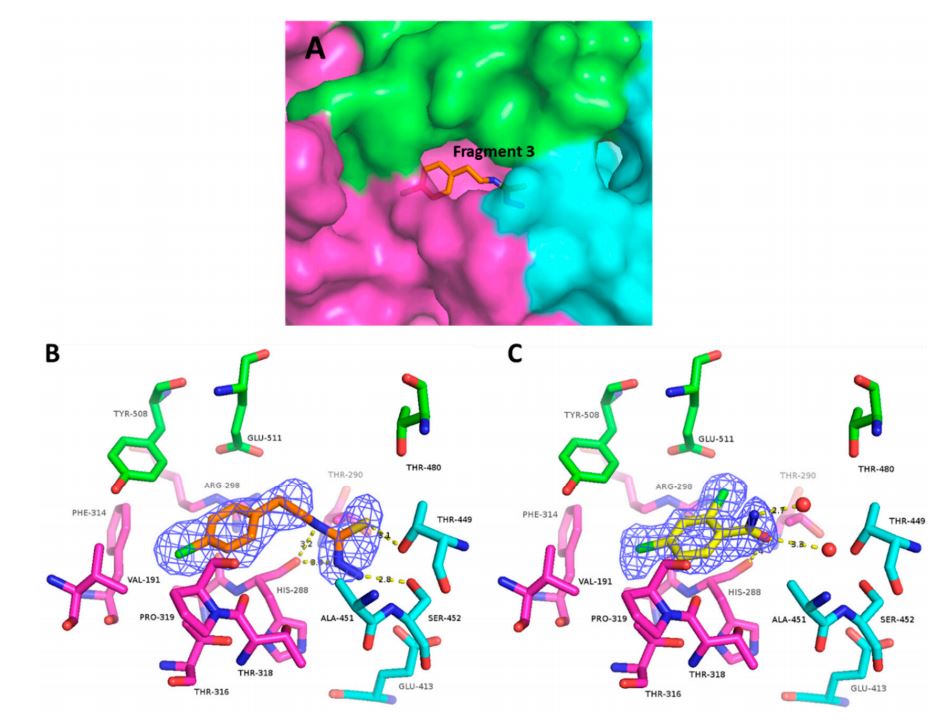 Fig.2 (A) Overall view of the binding pocket with fragment 3, (B) Electron density map and H-bonds made by fragment 3, (C) Electron density and H-bonds made by fragment 4. (Munawar, A.; et al. 2018)
Fig.2 (A) Overall view of the binding pocket with fragment 3, (B) Electron density map and H-bonds made by fragment 3, (C) Electron density and H-bonds made by fragment 4. (Munawar, A.; et al. 2018)Structure characterization
X-ray crystallography provides a complete three dimensional readout of the interaction between the small molecule and macromolecular target and reveals the structure and function of many biological molecules.
Identification of the binding sites and modes
X-ray crystallography can identify multiple active binding sites and help to for understand the binding mode of ligands, thus facilitating the subsequent modification of the fragment.
Our Services of X-Ray Crystallography
Crystal Soaking
Aiming to obtain complex crystals, we provide crystal soaking services in which our experts soak the liand in the native target protein crystal or replace the ligand previously bound to the target with the selected ligand.
Co-crystallization
With professional experts and rich experiences in crystal engineering, our teams can select the suitable crystallization conditions to meet different sizes or the chemical properties of the ligand. We also have abilities in performing high-standard protein-ligand co-crystallization and help to save your time and resources in crystals optimization.
Hit compounds validation
We can directly validate the hit compounds obtained through the crystal soaking and co-crystallization process.
Lead compound optimization
Our scientists can obtain a large amount of target-ligand complex structures in a short time and identify the binding mode of a fragment with their structural biology technique, offering the guidance in the optimization of the lead compound.
Our Advantages of X-Ray Crystallography Services
- Our X-ray crystallography platform can provide rapid access to detailed structural data about drug targets and bound ligands, giving important information to guide the identification and optimization of lead compounds.
- We offer high-throughput crystallography techniques to reduce the false positives and evaluate the validity with our automated and robotic equipment.
Equipment
Rapid and efficient protein crystal screening: mosquito® Xtal3.
Monitor and documentation of crystals:5 MP HD Microscope Camera Leica MC170 HD.
Reference
- Ke, T. W.; et al. A convolutional neural network-based screening tool for X-ray serial crystallography. Journal of Synchrotron Radiation. 2018, 25(3): 655-670.
- Munawar, A.; et al. Discovery of Novel Druggable Sites on Zika Virus NS3 Helicase Using X-ray Crystallography-Based Fragment Screening. International Journal of Molecular Sciences. 2018, 19(11).
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

