Homology Modeling
Working of Homology Modeling
As a most widely used computational methodology, this method provides various benefits due to its several advantages such as simplicity, reliability, and relatively high accuracy. There is a principle that when the sequence of two proteins have a high enough similarity, they tend to present similar three-dimensional structures. Therefore, homology modeling works on the basis of this principle and it is feasible to obtain a protein by copying the sequences of a protein whose structure is already known and identified. Scientists use this technique to establish the 3D model of a target protein and predict its structure.
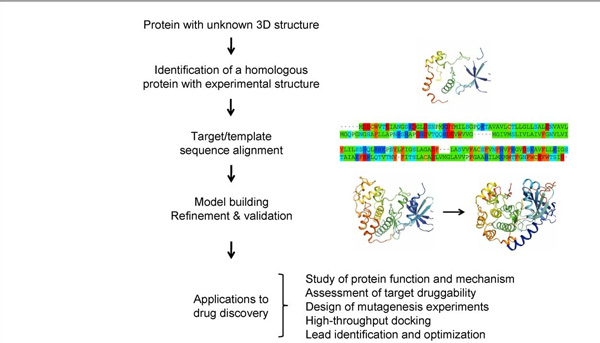 Fig.1 Outline of the homology modeling process and its applications in drug discovery. (Cavasotto, C. N.; Phatak, S. S. 2009)
Fig.1 Outline of the homology modeling process and its applications in drug discovery. (Cavasotto, C. N.; Phatak, S. S. 2009)Applications of Homology Modeling
Rational design and optimize ligands.
Docking of macromolecules prediction of protein partners.
Designing and validating proteins with better stability and new functions.
Provided as a useful analysis tool to assess the structural function of an unknown protein, providing guidance on the further experimental process.
Improving the structures obtained by other biophysical techniques such as NMR or X-ray crystallography.
Our Services of Homology Modeling
Query sequences
We firstly apply a protein with known structure (template) to conduct sequences searching campaigns to identify the primary target protein.
Alignment
In order to obtain the optimal framework of homologous proteins, our teams then align the query sequence with the candidate template and use diverse validated algorithms to correct the result and improve the quality of alignments.
3D structure construction
We perform rigid body assembly in which rigid bodies from aligned sequences, core region, loops, and side chains to obtain the structure of your target protein.
Model optimization
Addition and optimization of side chain atoms and loops are achieved with the use of skeleton copy, side chain construction, missing residues complement and loop modeling.
Model validation
Our teams apply some validation methods (including energy minimization procedure) to correct the structural irregularities.
Model evaluation
We assess the validated model to make sure that the final structure is consistent with each physicochemical property of the template.
Our Advantages of Homology Modeling Services
We are capable of delivering accurate sequence alignment with manual corrections.
BOC Sciences offers high-quality protein modeling services to help our customers better understand the 'structural-functional' relationship of target proteins.
Our well-built homologous models can be applied for studying the interaction patterns between the target and the drug, and other potential therapeutic fields.
Reference
Cavasotto, C. N .; Phatak, S. S. Homology modeling in drug discovery: current trends and applications. Drug Discovery Today, 2009, 14(13-14):676-683.
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

