Protein Nucleic Acid Docking
As a powerful tool in learning the protein-DNA/RNA interactions, protein-nucleic acid docking offers a deep insight into various essential biological processes which includes RNA transcription, translation, splicing and degradation, or DNA replication, repair, and protein synthesis. This method is applied to identify amino acids or nucleotide residues through modeling protein-nucleic acid complex structures.
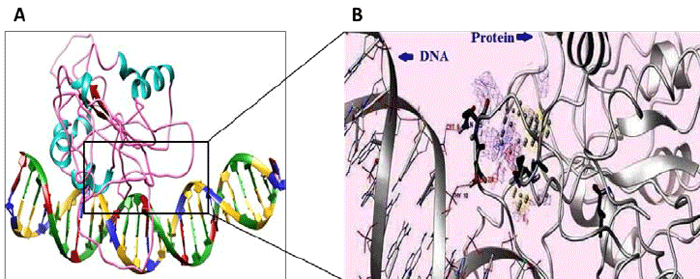 Fig.1 (A) Protein-DNA docking model and DNA segment. (B) Enlarged molecular view of rectangle. (Pandey, D. M. 2015)
Fig.1 (A) Protein-DNA docking model and DNA segment. (B) Enlarged molecular view of rectangle. (Pandey, D. M. 2015)Our Services of Protein Nucleic Acid Docking
Docking
We carry out a rigid body searching to generate protein-nucleic acid complex structures.
Scoring and ranking
Our teams use diverse statistical methods to score and rank the resulting models.
Optimization
The best-scored protein-RNA/DNA structures are then clustered in this step, and the largest ones are selected for the subsequent optimization of structure.
Our Advantages of Protein Nucleic Acid Docking Services
We can apply user-defined restraints to limit the search space, delivering a rapid and satisfying outcome of searching.
We have developed a diversity of computational docking approaches to search all possible binding patterns in the translational and rotational space between nucleic acids and their target proteins.
Our docking capabilities are also able to support docking of some hybrid DNA-RNA molecules, offering our customers comprehensive docking services.
Our Capabilities of Protein Nucleic Acid Docking Services
In the scoring and ranking process, our statistical strength can provide our customers reliable assessment of protein-nucleic acid interactions, which helps you better understand these complex structures.
We have abilities in selecting the highly populated clusters of low-energy conformations in the clustering campaigns.
We validate and deliver our final predictions based on various filter factors, such as blocking residues, binding site residues, pairwise distance restraints.
At BOC Sciences, the flexible refinement and modification of amino acids and DNA/RNA molecules are achieved to meet your demand.
Reference
Pandey, D. M.Molecular beacon probe based promoter motifs validation in anoxia responsive differentially expressed genes and their in silico interaction studies with AP2/EREBP TF in rice (Oryza Sativa L.). International Journal of Pharmacy & Pharmaceutical Sciences. 2015, 7(3): 123-130.
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

