ADMET Modeling and Prediction
Prediction of ADMET (absorption, distribution, metabolism, excretion, and toxicity) is an essential step in drug design and drug screening. ADME gives accurate descriptions of the disposition of a chemical compound within an organism. ADME/Tox prediction has been applied to investigate the drug levels and kinetics of candidates when it exposures to the tissues. Scientists can modify the pharmacological activity of the compound by predicting these characteristics rapidly.
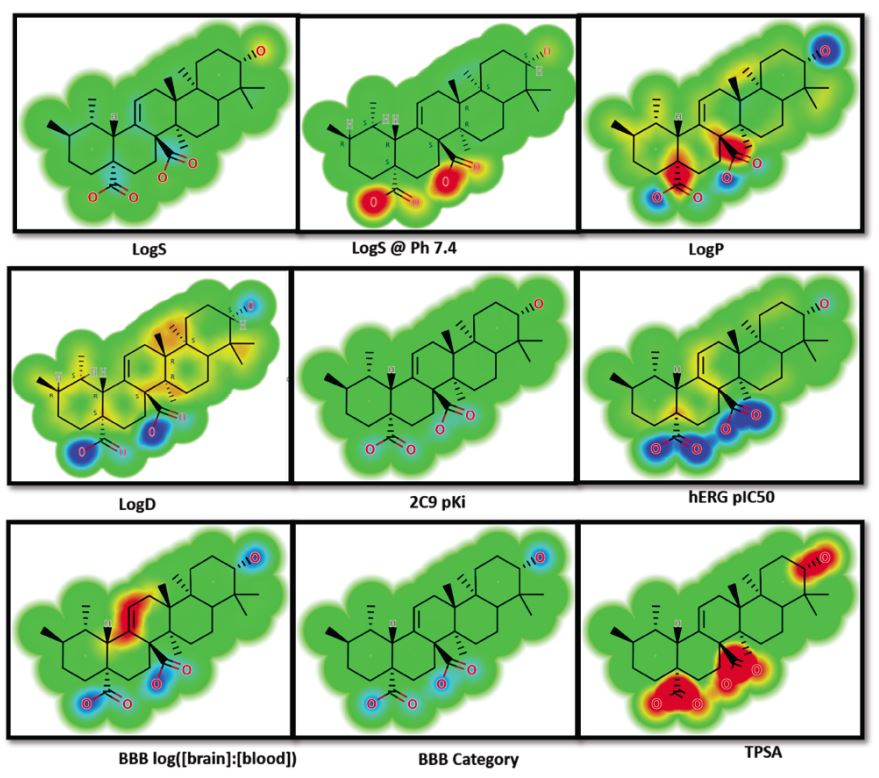 Fig.1 ADME influencing structural features in quinovic acid. (Rajeev, K. S.; Bairong, S. 2021)
Fig.1 ADME influencing structural features in quinovic acid. (Rajeev, K. S.; Bairong, S. 2021)Prediction Workflow At BOC Sciences
Data collection
We collect data from various public databases through manually filtering and multi-step processing.
Data preparation
We perform pretreatments by removing compounds without detailed and reliable information of ADME/Tox properties.
Descriptor Selection
Our scientists develop a validated regression model by filtering descriptors with invalid information or intrusive.
Modeling
At BOC Sciences, two kinds of computational approaches are applied: Molecular modelling and data modelling.
Evaluation
We generate a set of prediction models with the application of validation approaches and crucial parameters to evaluate various ADMET properties.
Our Abilities of ADMET Modeling and Prediction
- We can reduce the random error when fluctuations of compounds values in a reasonable limit wash molecules by molecular operating environment software.
- Multiple widely used molecular descriptors are available for further model building including constitution, topology, connectivity, E-state, Kappa, basak, burden, autocorrelation, charge property, MOE-type descriptors, etc.
- Different modeling algorithms are applied to develop regression or classification models for ADME/T related properties:
- We are capable of evaluating various predicted properties including:
- Our computational modeling capabilities
RF, SVM, RP and PLS are used for regression model building.
RF, SVM, NB and DT are applied to build those classification models.
Some basic physiochemical properties: Aqueous (water) solubility, boiling point, logD, logP, LogD7.4, pKa, MW, number of hydrogen bond donors/acceptors and rotatable bonds, molar refractivity/volume, polarizability.
Absorption: Intestinal absorption, Caco-2 permeability, Pgp-Inhibitor/substrate, HIA, oral bioavailability
Distribution: Plasma protein binding, Vd, P-gp, AUC, PPB, VD, BBB.
Metabolism: Half-time, CYP-Inhibitor/substrate, drug-drug interaction.
Excretion: Urinary excretion, clearance, DILI
Toxicity: Acute/aquatic toxicity, genotoxicity, hERG Inhibition, H-HT, Ames, SkinSen.
Molecular modelling: We apply protein modeling when the 3D structure of proteins are known as well as homology modelling and pharmacohphore models while the protein structure is not available.
Data modelling: At BOC Sciences, we introduce QSAR and QSPR approaches to investigate a variety of biological and physicochemical data by searching for correlations between a given property and a set of molecular of structural descriptors of the molecules.
Our Advantages of ADMET Modeling and Prediction
- We offer accurate and liable predicted ADME-Tox data to assist your virtual screening.
- Our groups are confident in the estimation for classification models.
- At our ADME/Tox prediction platforms, other properties such as bioavailability, safety, as well as activity, are able to be investigated in parallel.
- We have extensive experience in pharmacophore modeling and QSAR modeling, delivering high-quality customized models.
- We are committed in the development of more effective ADME-Tox models utilizing large and well-validated datasets.
Reference
- Rajeev, K. S.; Bairong, S.In Silico ADMET Evaluation of Natural DPP-IV Inhibitors for Rational Drug Design against Diabetes. Current Drug Metabolism. 2020, 21.
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

