Ligand Based Virtual Screening
Ligand-based virtual screening method is developed based on the 'similarity principle' which means similar molecules tend to exhibit approximative biological properties. In the LBVS, a group of ligands with diverse structures (which exhibit beneficial effect in a reference ligand) can be obtained by similarity or pharmacophore searching and their receptor models are built with the use of the collective information of the group of ligands.
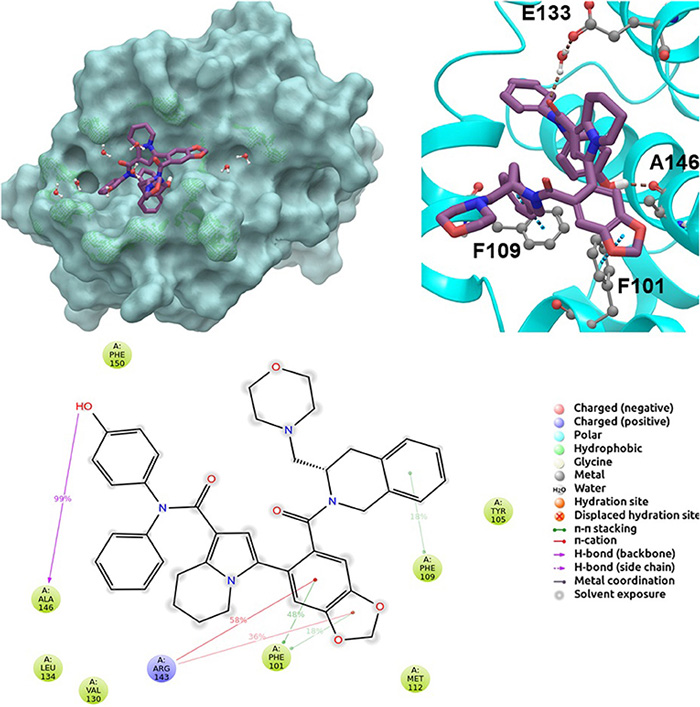 Fig.1 2D and 3D ligand interactions diagrams of selected molecule S55746 at the binding pocket. (Tutumlu, G.; et al. 2020)
Fig.1 2D and 3D ligand interactions diagrams of selected molecule S55746 at the binding pocket. (Tutumlu, G.; et al. 2020)Our Services of Ligand Based Virtual Screening
Similarity searching
BOC Sciences has collected a huge amount of fingerprints which give accurate and detailed description of the chemical characteristics (physicochemical properties, 2D properties, and 3D properties) of ligands.
We can perform a rapid similarity searching of diverse active compounds with the use of data integration and machine learning.
Quantitative structure-activity relationship (QSAR) modeling
We provide QSAR analysis to support the optimization of hits and lead compounds with our professional mathematical knowledge. Our experts have abilities and experience in generating QSAR models using diverse molecular descriptors and multidimensional fingerprints, giving reliable prediction of the biological properties of your new compounds.
Pharmacophore modeling
A pharmacophore model is used to virtual screen of new ligands that match the pharmacophore features. It identifies important molecular recognition and gives characterization of molecules on diagrammatic 3D level. At BOC Sciences, we offer ligand-based pharmacophore modeling and structure-based pharmacophore modeling methods to build pharmacophore models according to your specific need.
Advantages of Our Ligand-Based Virtual Screening Services
- We provide a diversity of compound databases containing over 10 million purchasable compounds to support your screening campaigns.
- Our computer-aided drug design platforms provide high-performance computer cluster which equipped with advanced servers and screening algorithms for high-throughput and large-scale LBVS.
- At BOC Sciences, our in silico and experimental teams have rich exiperience in the development of pharmacophore, similarity methods, 2D/3D QSAR models and machine learning.
- We are capable of integrating diverse cost-effective methods to deliver the most promising hits.
Reference
- Tutumlu, G.; et al.Integrating Ligand and Target-Driven Based Virtual Screening Approaches With in vitro Human Cell Line Models and Time-Resolved Fluorescence Resonance Energy Transfer Assay to Identify Novel Hit Compounds Against BCL-2. Frontiers in Chemistry. 2020, 8: 167.
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

