De Novo Drug Design
De novo drug design is a computational method which is utilized in the fields of small molecular design and lead discovery. The goal of this artificial intelligence-driven technology is to generate new molecular structures with desired pharmacological properties. Medicinal chemists design a new molecule utilizing the 3D structure of the receptor, and the process involves structure determination of the lead target and the design of lead modifications using modeling tools.
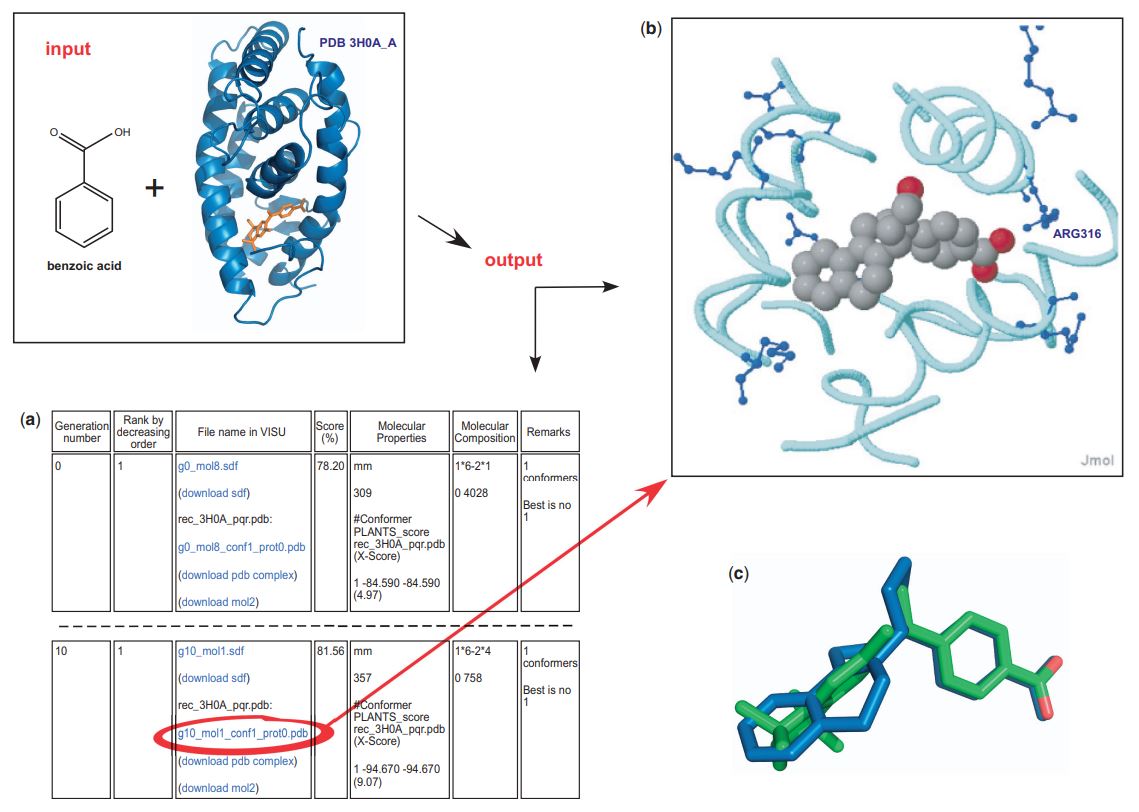 Fig.1 A de novo drug design module. (Dominique, D. 2010)
Fig.1 A de novo drug design module. (Dominique, D. 2010)Applications of De Novo Drug Design
- Explore a broader chemical space.
- Design a novel set of classes of chemical compounds with similar substituent to the target by using a template or scaffold.
- Discovery compounds that enable to constitute novel intellectual property.
According to each specific project requirement of our clients, we offer structure-based de novo drug design (SBDND) and ligand-based de novo drug design (LBDND).
Our Services of SBDND
Structure construction
We firstly define the active site of the receptor through studying various receptor-ligand interactions.
Optimization
We then optimize the structure by using a force-field to minimize most of the functional groups.
Evaluation
The final evaluation of the molecule class is performed by 3D protein-ligand docking as well as the calculation of the candidate molecule’s free binding energy using scoring functions.
Validation
We provide the service of examining the searching space.
Our Services of LBDND
Structure construction
We use the data of known active binders and establish a ligand pharmacophore model to design novel structures.
Optimization
We apply a quantitative structure-activity relationship model and ligand-based similarity measure to evaluate the quality of the fitness of a molecule.
Examination
We perform estimation of the search space or difficulty of optimization.
Computational Methods at BOC Sciences
Evolutionary algorithms
Site point connection method: LUDI.
Fragment connection methods: SPLICE, NEW LEAD, PRO-LIGAND.
Sequential build up methods: LEGEND, GROW, and SPROUT.
Random connection and disconnection methods: CONCEPTS, CONCERTS, MCDNLG.
Benchmark fingerprint descriptors
AtomPairs fingerprints, Morgan fingerprints, RDKit fingerprints, MACCS keys.
Our Capabilities of De Novo Drug Design
- We have developed multiple computerized structural design programs in the structure-base design and ligand-based design.
- Our scoring functions abilities involve: Force fields, empirical scoring functions, and knowledge-based scoring functions.
- The final validation process is carried out on the basis of depth-or breadth-first search, evolutionary algorithms and exhaustive structure enumeration.
- Our teams can optimize crucial parameters of QSAR/QSPR models to offer high-quality structure-activity relationship analysis.
- At BOC Sciences, various target prediction methods, molecular shape and partial charge descriptors are available to determine the similarity of the designed compounds to known bioactive chemicals.
Reference
- Dominique, D.e-LEA3D: a computational-aided drug design web server. Nucleic Acids Research. 2010, 38: 615-621.
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

