Pharmacophore Modeling
What is Pharmacophore Modeling?
Each type of atom or group in a compound exhibiting chemical properties can be recognized and simplified into a pharmacophore feature by the developed model. A pharmacophore shows the spatial arrangements of molecular features and confirms the optimal molecular interactions.
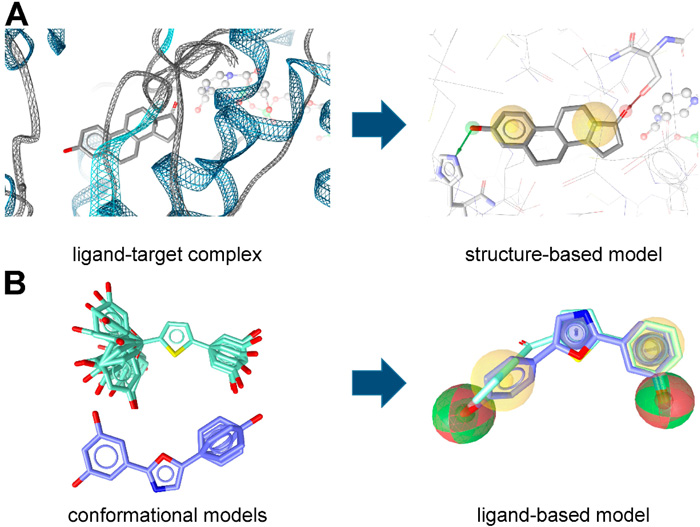 Fig.1 Two kinds of pharmacophore models. (Teresa, K.; et al. 2015)
Fig.1 Two kinds of pharmacophore models. (Teresa, K.; et al. 2015)Methodologies of Pharmacophore Modeling
Based on the detailed information of your project, we introduce two basic approaches to establish pharmacophore models:
- When the structure of target is unknown, our scientists perform ligand-based pharmacophore modeling method in which we extract common chemical characteristics from 3D structures of multiple reference ligands and conduct the searching of pharmacophores that would match the active compound.
- The structure-based pharmacophore modeling is conducted based on target-ligand complexes and generates the selected chemical features and sterical relationships. We build this model by detecting the possible molecular interaction sites.
Applications of Pharmacophore Modeling
- We guide the molecular docking and simulation, improving the outcomes of your virtual screening with our flexible solutions.
- The identification of the interactions of drugs with drug-metabolizing enzymes can be achieved through pharmacophore methods, which means you can conduct ADME studies and toxicity prediction.
- Our collections of diverse models can identify the drug-like molecules that the model fits well and enable to identify the target with a unknown activity.
Our Services of Pharmacophore Modeling
- Our scientists are capable of identifying the key pharmacophore features of bioactive molecules and build quantitative structure-activity relationships (QSAR) models based on them.
- We have designed modern ligand alignment algorithms and machine learning algorithms with accurate pharmacophore fingerprints.
- We develop a pharmacophore method to perform virtual screening and search for new scaffold-based compounds with similar biological activity. And we also provide the subsequent services for synthesizing novel active molecules.
- We optimize the outcome of docking simulations by removing compounds that fail to bind according to the pharmacophore query.
- We develop 3D pharmacophore models in the molecular dynamics computational studies to modulate bioactivity by presenting a physiologically relevant dynamic interaction pattern.
- We perform ADME-tox prediction by matching the chemical groups of target molecules to drug molecules with known ADME-tox profiles.
- We can perform target-fishing and provide prediction of possible side effects or off-targets, improving your outcome of target investigation.
- Our experts use pharmacophore knowledge to give additional support for de novo drug design to assess the drug-likeness and bioactivity of the designed compounds.
Our Advantages of Pharmacophore Modeling Services
- BOC Sciences has multiple strategies to perform both ligand-based and structure-based pharmacophore methods depending on the type of your experiments.
- We are capable of offering 2D and 3D pharmacophore model buildings development.
- Our groups can conduct virtual screening based on pharmacophore approaches with advanced and high performance computing software.
- We have professional competence in the determination of the structure of targets and target-ligand complexes.
Reference
- Teresa, K.; et al. Pharmacophore Models and Pharmacophore-Based Virtual Screening: Concepts and Applications Exemplified on Hydroxysteroid Dehydrogenases. Molecules. 2015, 20(12).
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

