Virtual Screening (VS)
Virtual screening is a powerful alternative approach to discover new drugs. The aim of this technology is to screen out potential leads from a huge number of compounds for further experimental activity evaluation based on drug design theory with the application of advanced computer tools and professional software. Scientists use VS approach to measure the presence or absence of specific substructures, match certain calculated molecular properties, and fit potential ligand molecules into the target receptor site.
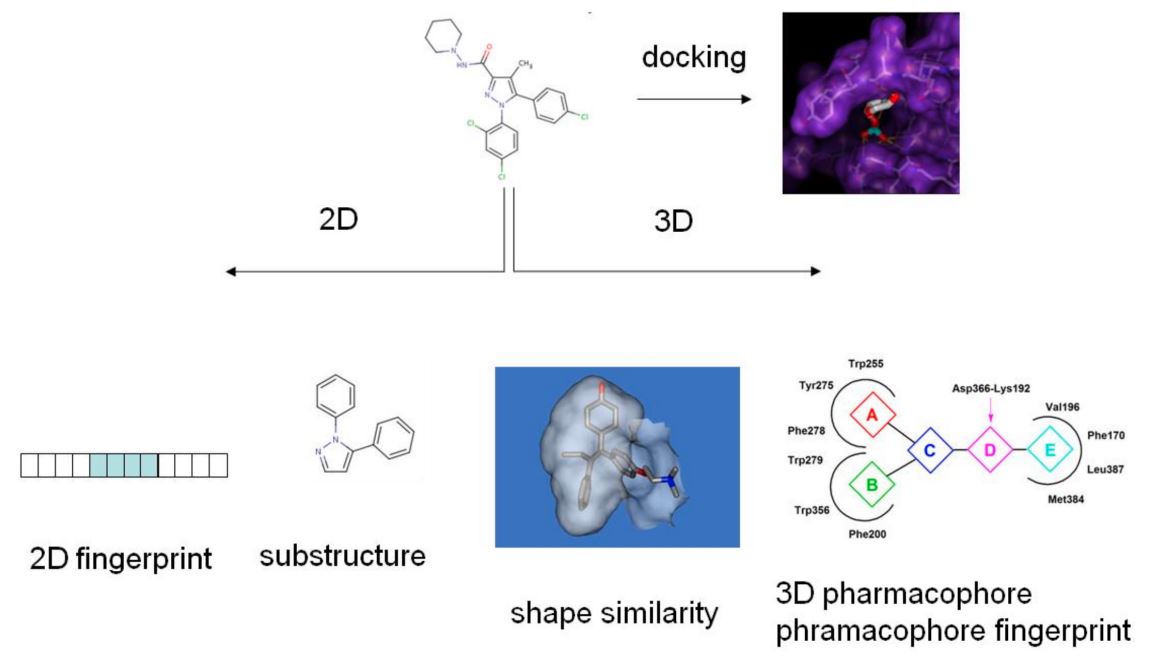 Fig.1 Schematic representation of virtual screening approaches. (Polgar, T.; Keseru, G. M. 2011)
Fig.1 Schematic representation of virtual screening approaches. (Polgar, T.; Keseru, G. M. 2011)VS Process
Screening of databases
Design the virtual compound libraries containing various drug-like compounds to discover novel scaffolds.
Use either structure-based approaches or ligand-based approaches combined the application of machine learning and other technologies.
Further filtering
Evaluate the ADME properties and toxicity to re-rank the compounds.
Compound selection & in vitro assay
Merits of VS
Cost-effective
VS reduces the number of compounds required to be synthesized or purchased.
Higher hit rate
VS expands the chemical space by constructing the virtual libraries of compounds.
Shorten the research cycle
Virtual Screening Services
Our groups offer high-quality virtual screening services using well-designed database and diverse computational tools. Several screening approaches available at BOC Sciences including but not limited to:
Virtual compound libraries
Virtual compound libraries are applied in searching of small molecules in order to identify promising structures which are most likely to bind to a drug target. BOC Sciences has a wealth of database resources and high-performance computer servers, capable of providing professional virtual screening services.
Hybrid methods
As an essential element of virtual screening method, hybrid approach provides further insights into the protein binding by combining structure-based and ligand-based drug design strategy. Our experts offer different hybrid techniques that integrate the structure information and ligand similarity of targets, predicting the biological activities and new drugs design.
Machine learning
Machine learning enables to produce abundant and high-quality data for virtual screening. This method can be applied in identifying which category a compound belongs to, predicting a continuous-values attribute associated with a compound and grouping of similar molecules into sets automatically. We have established well-designed computational algorithms to perform both vector machine and binary kernel discrimination.
Structure-based virtual screening (SBVS)
SBVS method uses the 3D structure of the target to analyze its binding site and then evaluate the protein-ligand binding modes based on the scoring function. Our platforms provide high-quality structure-based virtual screening services combining homology modeling, molecular docking and molecular dynamics simulations, which can generate the binding conformation of target-ligand complexes and select optimal compound for subsequent biological activity test.
Ligand-based virtual screening (LBVS)
LBVS strategy focuses on analyzing structures and physicochemical properties of compounds and it is applied in the quantitative structure-activity relationship analysis and pharmacophore modeling to predict the bioactivity of novel compounds. BOC Sciences can utilize 2D or 3D chemical structures to perform screening of databases using different types of similarity measurements, searching for compounds that are active towards a target of interest.
De novo drug design
De novo drug design is a time-saved and cost-effective technology to generate novel molecular structures with desired pharmacological properties. Both structure-based de novo drug design and ligand-based de novo drug design are available at BOC Sciences, and our experts are committed to offering customized solutions to help you design and identify new lead compounds.
ADME/Tox prediction
Prediction of ADMET plays an important role in drug screening and design. This method evaluates ADME-Tox properties to determine whether a drug candidate is safe enough and has satisfying efficacy. We perform diverse modeling algorithms to assess various physicochemical properties of compounds, predicting the interaction between a drug and its target protein and whether the drug has a effect in human's bodies.
Our Advantages of Virtual Screening Services
- We are equipped with super high-performance computer server.
- At BOC Sciences, well-designed libraries and abundant database resources are available.
- Our professional molecular simulation and drug design teams provide support for the selection and identification of promising leads.
- Our comprehensive CADD platform can improve the performance of VS and support high-throughput virtual drug screening.
Reference
- Polgar, T.; Keseru, G. M. Integration of Virtual and High Throughput Screening in Lead Discovery Settings. Combinatorial Chemistry & High Throughput Screening. 2011, 14(10).
※ It should be noted that our service is only used for research.

One-stop
Drug Discovery Services
- Experienced and qualified scientists functioning as project managers or study director
- Independent quality unit assuring regulatory compliance
- Methods validated per ICH GLP/GMP guidelines
- Rigorous sample tracking and handling procedures to prevent mistakes
- Controlled laboratory environment to prevent a whole new level of success
Online Inquiry

